8.3 KiB
ggalt : Alternate/Extra 'Geoms', 'Stats' and 'Coords' for 'ggplot2'
A package containing additional/alternate 'geoms', 'coords' and 'stats' for use with the revamped (late 2015) version of ggplot2.
The first three forays into this brave, new ggplot2 world are splines! and being able to use the (much better) KernSmooth::bkde and KernSmooth::bkde2D for density plots and an initial port of the (still needing work) coord_proj.
NOTE
Until the new ggplot2 version is on CRAN, you'll need to install it from github via devtools::install_github("hrbrmstr/ggplot2"). Locally, I have goth ggalt and my ggplot2 in a "develment mode" install via devtools::dev_mode(). Since the new ggplot2 breaks many other packages (like plotly, CRAN ggthemes, ggmap and more), keeping it squirreled away in it's own area is a good idea until everyone catches up.
The following functions are implemented:
coord_proj: Likecoord_maponly better:-)geom_xspline: Connect control points/observations with an X-splinestat_xspline: Connect control points/observations with an X-splinegeom_bkde: Display a smooth density estimate (usesKernSmooth::bkde)stat_bkde: Display a smooth density estimate (usesKernSmooth::bkde)geom_bkde2d: Contours from a 2d density estimate. (usesKernSmooth::bkde2D)stat_bkde2d: Contours from a 2d density estimate. (usesKernSmooth::bkde2D)stat_ash: Compute and display a univariate averaged shifted histogram (polynomial kernel) (usesash::ash1/ash::bin1)
News
- Version 0.0.4.9000 released -
stat_ash - Version 0.0.3.9000 released -
coord_proj! (requires my github copy of ggplot2 for now) - Version 0.0.2.9005 released - cleanup before blog post
- Version 0.0.2.9002 released - working 2D density plots
- Version 0.0.2.9000 released
Installation
# you'll want to see the vignettes, trust me
devtools::install_github("hadley/ggplot2", build_vignettes=TRUE)
devtools::install_github("hrbrmstr/ggalt")
Usage
library(ggplot2)
library(gridExtra)
library(ggalt)
# current verison
packageVersion("ggalt")
#> [1] '0.0.4.9000'
set.seed(1492)
dat <- data.frame(x=c(1:10, 1:10, 1:10),
y=c(sample(15:30, 10), 2*sample(15:30, 10), 3*sample(15:30, 10)),
group=factor(c(rep(1, 10), rep(2, 10), rep(3, 10)))
)
Splines!
ggplot(dat, aes(x, y, group=group, color=group)) +
geom_point() +
geom_line()

ggplot(dat, aes(x, y, group=group, color=factor(group))) +
geom_point() +
geom_line() +
geom_smooth(se=FALSE, linetype="dashed", size=0.5)

ggplot(dat, aes(x, y, group=group, color=factor(group))) +
geom_point(color="black") +
geom_smooth(se=FALSE, linetype="dashed", size=0.5) +
geom_xspline(size=0.5)

ggplot(dat, aes(x, y, group=group, color=factor(group))) +
geom_point(color="black") +
geom_smooth(se=FALSE, linetype="dashed", size=0.5) +
geom_xspline(spline_shape=-0.4, size=0.5)

ggplot(dat, aes(x, y, group=group, color=factor(group))) +
geom_point(color="black") +
geom_smooth(se=FALSE, linetype="dashed", size=0.5) +
geom_xspline(spline_shape=0.4, size=0.5)
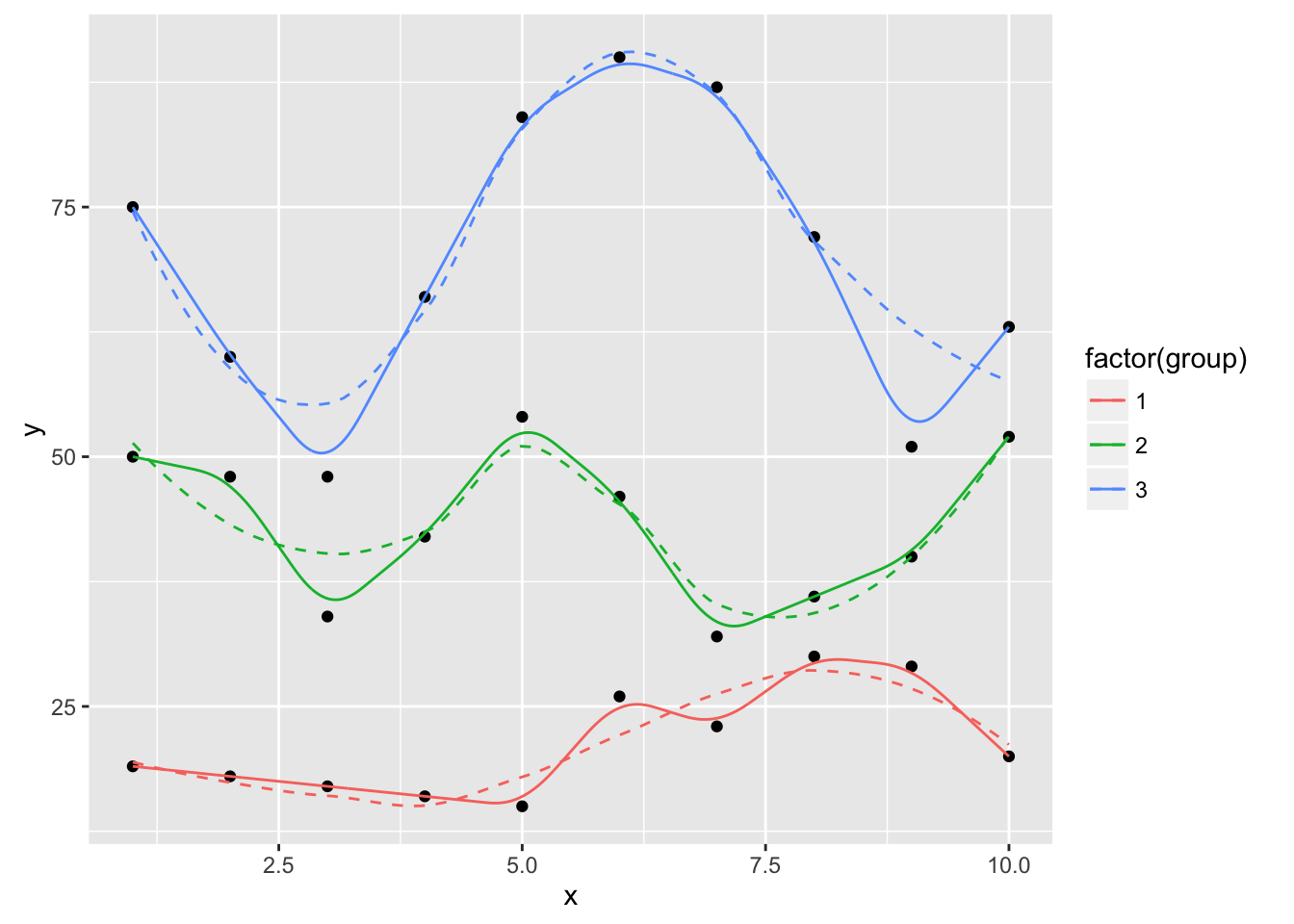
ggplot(dat, aes(x, y, group=group, color=factor(group))) +
geom_point(color="black") +
geom_smooth(se=FALSE, linetype="dashed", size=0.5) +
geom_xspline(spline_shape=1, size=0.5)

ggplot(dat, aes(x, y, group=group, color=factor(group))) +
geom_point(color="black") +
geom_smooth(se=FALSE, linetype="dashed", size=0.5) +
geom_xspline(spline_shape=0, size=0.5)

ggplot(dat, aes(x, y, group=group, color=factor(group))) +
geom_point(color="black") +
geom_smooth(se=FALSE, linetype="dashed", size=0.5) +
geom_xspline(spline_shape=-1, size=0.5)

Alternate (better) density plots
# bkde
data(geyser, package="MASS")
ggplot(geyser, aes(x=duration)) +
stat_bkde(alpha=1/2)
#> Bandwidth not specified. Using '0.14', via KernSmooth::dpik.

ggplot(geyser, aes(x=duration)) +
geom_bkde(alpha=1/2)
#> Bandwidth not specified. Using '0.14', via KernSmooth::dpik.
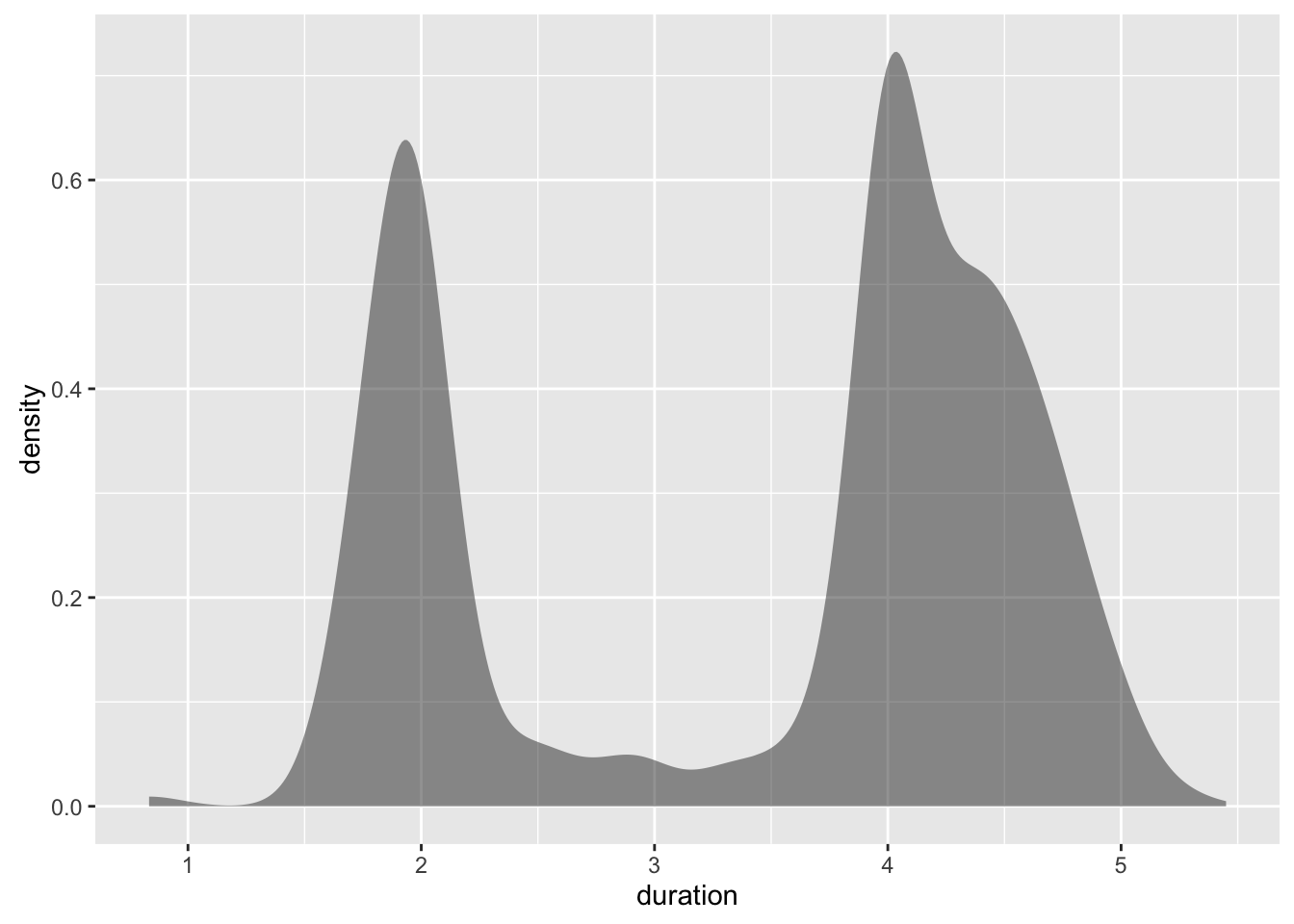
ggplot(geyser, aes(x=duration)) +
stat_bkde(bandwidth=0.25)

ggplot(geyser, aes(x=duration)) +
geom_bkde(bandwidth=0.25)

set.seed(1492)
dat <- data.frame(cond = factor(rep(c("A","B"), each=200)),
rating = c(rnorm(200),rnorm(200, mean=.8)))
ggplot(dat, aes(x=rating, color=cond)) + geom_bkde(alpha=0)
#> Bandwidth not specified. Using '0.36', via KernSmooth::dpik.
#> Bandwidth not specified. Using '0.31', via KernSmooth::dpik.

ggplot(dat, aes(x=rating, fill=cond)) + geom_bkde(alpha=0.3)
#> Bandwidth not specified. Using '0.36', via KernSmooth::dpik.
#> Bandwidth not specified. Using '0.31', via KernSmooth::dpik.

# ash
set.seed(1492)
dat <- data.frame(x=rnorm(100))
grid.arrange(ggplot(dat, aes(x)) + stat_ash(),
ggplot(dat, aes(x)) + stat_bkde(),
ggplot(dat, aes(x)) + stat_density(),
nrow=3)
#> Estimate nonzero outside interval ab.
#> Bandwidth not specified. Using '0.43', via KernSmooth::dpik.

cols <- RColorBrewer::brewer.pal(3, "Dark2")
ggplot(dat, aes(x)) +
stat_ash(alpha=1/2, fill=cols[3]) +
stat_bkde(alpha=1/2, fill=cols[2]) +
stat_density(alpha=1/2, fill=cols[1]) +
geom_rug() +
labs(x=NULL, y="density/estimate") +
scale_x_continuous(expand=c(0,0)) +
theme_bw() +
theme(panel.grid=element_blank()) +
theme(panel.border=element_blank())
#> Estimate nonzero outside interval ab.
#> Bandwidth not specified. Using '0.43', via KernSmooth::dpik.

Alternate 2D density plots
geyser_dat <- data.frame(x=geyser$duration, y=geyser$waiting)
ggplot(geyser_dat, aes(x, y)) +
geom_point() +
geom_bkde2d(bandwidth=c(0.7, 7))

ggplot(geyser_dat, aes(x, y)) +
geom_point() +
stat_bkde2d(bandwidth=c(0.7, 7))

coord_proj LIVES! (still needs work)
# devtools::install_github("hrbrmstr/ggplot2")
world <- map_data("world")
#>
#> # ATTENTION: maps v3.0 has an updated 'world' map. #
#> # Many country borders and names have changed since 1990. #
#> # Type '?world' or 'news(package="maps")'. See README_v3. #
world <- world[world$region != "Antarctica",]
gg <- ggplot()
gg <- gg + geom_map(data=world, map=world,
aes(x=long, y=lat, map_id=region))
gg <- gg + coord_proj("+proj=wintri")
gg

Test Results
library(ggalt)
library(testthat)
date()
#> [1] "Sat Oct 31 10:07:15 2015"
test_dir("tests/")
#> testthat results ========================================================================================================
#> OK: 0 SKIPPED: 0 FAILED: 0
Code of Conduct
Please note that this project is released with a Contributor Code of Conduct. By participating in this project you agree to abide by its terms.